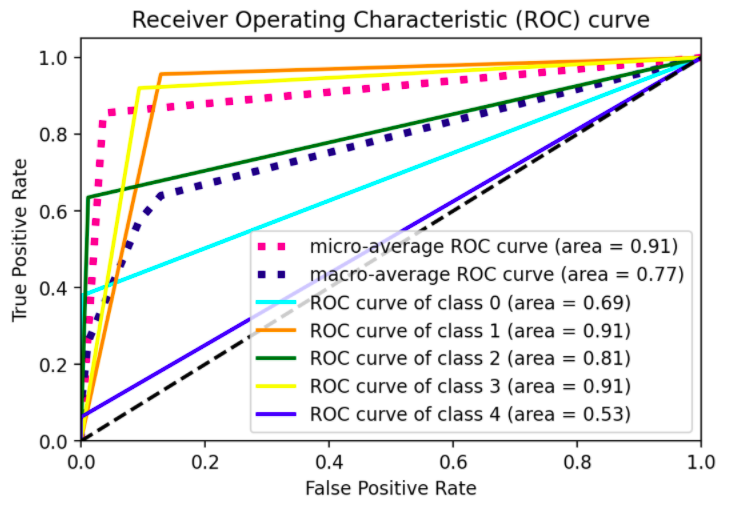
Plotting the ROC curve for a multiclass problem
I am trying to apply the idea of sklearn ROC extension to multiclass to my dataset. My per-class ROC curve looks find of a straight line each, unline the sklearn's example showing curve's fluctuating.
I give an MWE below to show what I mean:
# all imports
import numpy as np
import matplotlib.pyplot as plt
from itertools import cycle
from sklearn import svm, datasets
from sklearn.metrics import roc_curve, auc
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import label_binarize
from sklearn.datasets import make_classification
from sklearn.ensemble import RandomForestClassifier
# dummy dataset
X, y = make_classification(10000, n_classes=5, n_informative=10, weights=[.04, .4, .12, .5, .04])
train, test, ytrain, ytest = train_test_split(X, y, test_size=.3, random_state=42)
# random forest model
model = RandomForestClassifier()
model.fit(train, ytrain)
yhat = model.predict(test)
The following function then plots the ROC curve:
def plot_roc_curve(y_test, y_pred):
n_classes = len(np.unique(y_test))
y_test = label_binarize(y_test, classes=np.arange(n_classes))
y_pred = label_binarize(y_pred, classes=np.arange(n_classes))
# Compute ROC curve and ROC area for each class
fpr = dict()
tpr = dict()
roc_auc = dict()
for i in range(n_classes):
fpr[i], tpr[i], _ = roc_curve(y_test[:, i], y_pred[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
# Compute micro-average ROC curve and ROC area
fpr["micro"], tpr["micro"], _ = roc_curve(y_test.ravel(), y_pred.ravel())
roc_auc["micro"] = auc(fpr["micro"], tpr["micro"])
# First aggregate all false positive rates
all_fpr = np.unique(np.concatenate([fpr[i] for i in range(n_classes)]))
# Then interpolate all ROC curves at this points
mean_tpr = np.zeros_like(all_fpr)
for i in range(n_classes):
mean_tpr += np.interp(all_fpr, fpr[i], tpr[i])
# Finally average it and compute AUC
mean_tpr /= n_classes
fpr["macro"] = all_fpr
tpr["macro"] = mean_tpr
roc_auc["macro"] = auc(fpr["macro"], tpr["macro"])
# Plot all ROC curves
#plt.figure(figsize=(10,5))
plt.figure(dpi=600)
lw = 2
plt.plot(fpr["micro"], tpr["micro"],
label="micro-average ROC curve (area = {0:0.2f})".format(roc_auc["micro"]),
color="deeppink", linestyle=":", linewidth=4,)
plt.plot(fpr["macro"], tpr["macro"],
label="macro-average ROC curve (area = {0:0.2f})".format(roc_auc["macro"]),
color="navy", linestyle=":", linewidth=4,)
colors = cycle(["aqua", "darkorange", "darkgreen", "yellow", "blue"])
for i, color in zip(range(n_classes), colors):
plt.plot(fpr[i], tpr[i], color=color, lw=lw,
label="ROC curve of class {0} (area = {1:0.2f})".format(i, roc_auc[i]),)
plt.plot([0, 1], [0, 1], "k--", lw=lw)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel("False Positive Rate")
plt.ylabel("True Positive Rate")
plt.title("Receiver Operating Characteristic (ROC) curve")
plt.legend()
Output:
plot_roc_curve(ytest, yhat)
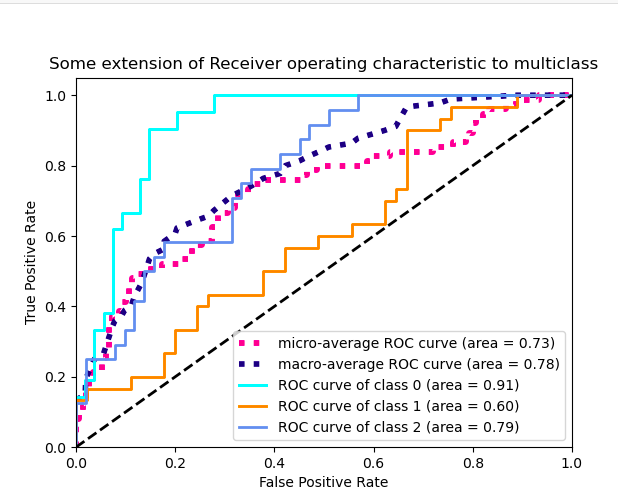
Kind of straight line bending once. I would like to see the model performance at different thresholds, not just one, a figure similar to sklearn's illustration for 3-classes shown below:
Point is that you're using
predict()rather thanpredict_proba()/decision_function()to define youry_hat. This means - considering that the threshold vector is defined by the number of distinct values iny_hat(see here for reference), that you'll have few thresholds per class only on whichtprandfprare computed (which in turn implies that your curves are evaluated at few points only).Indeed, consider what the doc says to pass to
y_scoresinroc_curve(), either prob estimates or decision values. In the example fromsklearn, decision values are used to compute the scores. Given that you're considering aRandomForestClassifier(), considering probability estimates in youry_hatshould be the way to go.What's the point then of label-binarizing the output? The standard definition for ROC is in terms of binary classification. To pass to a multiclass problem, you have to convert your problem into binary by using OneVsAll approach, so that you'll have
n_classnumber of ROC curves. (Observe, indeed, that asSVC()handles multiclass problems in a OvO fashion by default, in the example they had to force to use OvA by applyingOneVsRestClassifierconstructor; with aRandomForestClassifieryou don't have such problem as that's inherently multiclass, see here for reference). In these terms, once you switch topredict_proba()you'll see there's no much sense in label binarizing predictions.# all imports import numpy as np import matplotlib.pyplot as plt from itertools import cycle from sklearn import svm, datasets from sklearn.metrics import roc_curve, auc from sklearn.model_selection import train_test_split from sklearn.preprocessing import label_binarize from sklearn.datasets import make_classification from sklearn.ensemble import RandomForestClassifier # dummy dataset X, y = make_classification(10000, n_classes=5, n_informative=10, weights=[.04, .4, .12, .5, .04]) train, test, ytrain, ytest = train_test_split(X, y, test_size=.3, random_state=42) # random forest model model = RandomForestClassifier() model.fit(train, ytrain) yhat = model.predict_proba(test) def plot_roc_curve(y_test, y_pred): n_classes = len(np.unique(y_test)) y_test = label_binarize(y_test, classes=np.arange(n_classes)) # Compute ROC curve and ROC area for each class fpr = dict() tpr = dict() roc_auc = dict() thresholds = dict() for i in range(n_classes): fpr[i], tpr[i], thresholds[i] = roc_curve(y_test[:, i], y_pred[:, i], drop_intermediate=False) roc_auc[i] = auc(fpr[i], tpr[i]) # Compute micro-average ROC curve and ROC area fpr["micro"], tpr["micro"], _ = roc_curve(y_test.ravel(), y_pred.ravel()) roc_auc["micro"] = auc(fpr["micro"], tpr["micro"]) # First aggregate all false positive rates all_fpr = np.unique(np.concatenate([fpr[i] for i in range(n_classes)])) # Then interpolate all ROC curves at this points mean_tpr = np.zeros_like(all_fpr) for i in range(n_classes): mean_tpr += np.interp(all_fpr, fpr[i], tpr[i]) # Finally average it and compute AUC mean_tpr /= n_classes fpr["macro"] = all_fpr tpr["macro"] = mean_tpr roc_auc["macro"] = auc(fpr["macro"], tpr["macro"]) # Plot all ROC curves #plt.figure(figsize=(10,5)) plt.figure(dpi=600) lw = 2 plt.plot(fpr["micro"], tpr["micro"], label="micro-average ROC curve (area = {0:0.2f})".format(roc_auc["micro"]), color="deeppink", linestyle=":", linewidth=4,) plt.plot(fpr["macro"], tpr["macro"], label="macro-average ROC curve (area = {0:0.2f})".format(roc_auc["macro"]), color="navy", linestyle=":", linewidth=4,) colors = cycle(["aqua", "darkorange", "darkgreen", "yellow", "blue"]) for i, color in zip(range(n_classes), colors): plt.plot(fpr[i], tpr[i], color=color, lw=lw, label="ROC curve of class {0} (area = {1:0.2f})".format(i, roc_auc[i]),) plt.plot([0, 1], [0, 1], "k--", lw=lw) plt.xlim([0.0, 1.0]) plt.ylim([0.0, 1.05]) plt.xlabel("False Positive Rate") plt.ylabel("True Positive Rate") plt.title("Receiver Operating Characteristic (ROC) curve") plt.legend()
Eventually, consider that roc_curve() has also a drop_intermediate parameter meant for dropping suboptimal thresholds (it might be useful to know).
- Word Clouds using TabPy
- Extracting lines from two files where there is a match of value in specific columns
- Instantiate empty type-hinted list
- How to fix the GSException: "Container not found" even when the container exists?
- Find index of min value in a matrix
- Break statement in finally block swallows exception
- Does `anaconda` create a separate PYTHONPATH variable for each new environment?
- How can I replace a substring in a Python pathlib.Path?
- How to draw a line on an image in OpenCV?
- Match 2 strings then print both matches
- subprocess and exchanging json: How can I use read() on stdin non-blockingly?
- DNS request with scapy (Python)
- What is the difference between xpath() and findall()?
- if statement in Django template not working
- Extract header/footer from PDF (programmatically)
- How to create virtual env with Python 3?
- Type hinting / annotation (PEP 484) for numpy.ndarray
- Allowing resizing window pyGame
- Python Multiprocessing empty array
- Different / better approaches for calling python function from Java
- Calling Python Functions from Java (Without Jython, because it is too slow.)
- Best way to find the months between two dates
- Calling Java from Python
- Calling Java from Python
- Identifying outliers from openCV contours based on curvature?
- Unfolding a cartesian binned dataset into polar coordinates
- Why is facecolor argument in plot_surface() of matplotlib not working in python?
- Data class with argument optional only in init
- Repl.it Python 3 Short cut Comment/Uncomment a block
- How can I find where Python is installed on Windows?